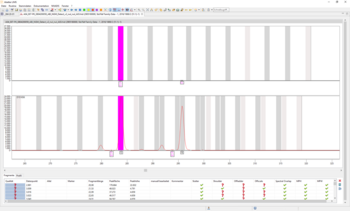
EasyRead technology and extension of electropherogram tools
With the update from July 2020, our customers will receive the improved algorithm for identification of sequencing and artificial artefacts. Especially improved for identification of pull-up artefacts, resulting from spectral overlaps. For easy identification and display of these artefacts, the electropherogram now includes highlighted bins. Thereby supporting the user visually in evaluating of alleles.
Our customers receive the update with the next product release, starting from July 2020.
Additional information to our EasyRead technology
Abetter LIMS uses the new EasyRead technology for evaluation of STR analysis data. This technology enables fast, robust and high-troughput evaluation of your sequencer data (fsa/hid). Even low template stains can easily be evaluated - without time-consuming changes to analysis methods or dependence on the analysis plattform. The EasyRead technology supports the 6-dye technology.
Size and allele calling
For fast and easy evaluation of STR data, the size and allele calling algorithms now include heuristic optimization for automatic peak detection via polynomial functions. Des Weiteren wird die Größenzuweisung durch eine umfangreiche Mustererkennung unterstützt. These improvements therefore allow for high troughput analysis.
Genetic Reference Data
Abetter LIMS supports the automatic analysis of autosomal and gonosomal markers with commercial multiplex or in-house kits. The analysis is based on an extensive genetic reference database, including most commecially available STR kits, size standards and markers. The user can add new data with multiple import options.
Graphic user interface
For the presentation and processing electropherograms, a unique graphic user interface has been developed. The user interface provides pixel perfect resolution, zoom functionalities, as well as various editing and evaluation tools for DNA mixtures. The user can also arrange windows to display electropherograms side by side or beneath each other for comparison of multiple profiles. To simplify analysis of multiple replicates, the marker centric view has been implemented.
Artefact detection
Abetter LIMS supports the complete process of raw data analysis with the help of highly developed algorithms for identification of artefacts. These artefacts include stutter peaks, shoulder peaks, off ladder peaks, off scale peaks and spectral overlap peaks. The artefact detection is included in the size and allele calling, but results can be adjusted by the users.